Beacon Aggregations for Data Summaries¶
WiP
Beacon aggregations are currently work in progress and may change without deprecation. They are not part of the official version 2.n specification and should not be implemented in production environments yet. Use with caution for testing, follow related PRs and provide feedback!
Overview and Use Cases¶
While the Beacon API provides different ways to discover and potentially retrieve
data in biomedical genomics resources, with version 2.n responses were limited to
global content (boolean or overall count of matched data and static collection information)
or full record level access which for most resources would not be possible in a
public context. Responses under the new aggregated granularity level allow to:
- provide granular data overwiews about the content of resources and their collections, e.g. numbers of samples with individual features or combinations of features
- profile query responses for multiple (single or intersected) parameters
Response Format¶
Aggregation are provided inside the responseAggregation property of the response and consist
of array of objects with the following structure:
- a required, ordered list of one or more
conceptsobjects, describing the parameters for which the aggregation is provided - summaries for the single or interssected concepts
distinctValuesCount: a count of the distinct values for single or intersected concepts and/oranyValueCount: a count for all records with existing values and/ordistribution: a distribution of all distinct values/combinations with the count of their occurrence
- an optional
scopeparameter to indicate the entity the results refer to (usually the current entry type but might be variable for collection and overview aggregations)
The following examples display different aggregation objects (which would be items in the responseAggregation array). Note that id values are for demonstration only and do not have a normative function.
Aggregations and Queries: For most of the example cases one can envision both a use in "data overview context" (e.g. to profile the content of a resource or collection) and in "query context" (e.g. to profile the response for a specific query).
Examples: Representation of Distinct Values Count distinctValuesCount¶
The most basic count - getting the number of samples:
{
"concepts": [
{"id": "biosampleCount", "property": "biosample.id"}
],
"distinctValuesCount": 258545
}
How many different diseases are represented in the data?
{
"concepts": [
{"id": "disease", "label": "Disease"}
],
"distinctValuesCount": 89
}
Example: Informative Values anyValueCount¶
How many individuals in the data have a follow-up time?
{
"scope": "individual",
"concepts": [
{"id": "followUpTime", "label": "Follow-up time"}
],
"anyValueCount": 1200
}
Example: Value Distribution distribution, Single Property¶
What is the distribution of diseases in the samples?
{
"scope": "biosample",
"concepts": [
{
"id": "sampleDiagnoses",
"label": "Diagnoses of selected carcinoma types",
"property": "biosample.histologicalDiagnosis.id"
}
],
"distribution": [
{
"conceptValues": [
{"id": "NCIT:C2919", "label": "Prostate Adenocarcinoma"}
],
"count": 426
},
{
"conceptValues": [
{"id": "NCIT:C4017", "label": "Breast Ductal Carcinoma"}
],
"count": 423
},
{
"conceptValues": [
{"id": "NCIT:C3512", "label": "Lung Adenocarcinoma"}
],
"count": 317
}
]
}
Example: Value Distribution distribution, Intersecting Concepts¶
What is the distribution of diseases in the samples, separately by sex? Please note:
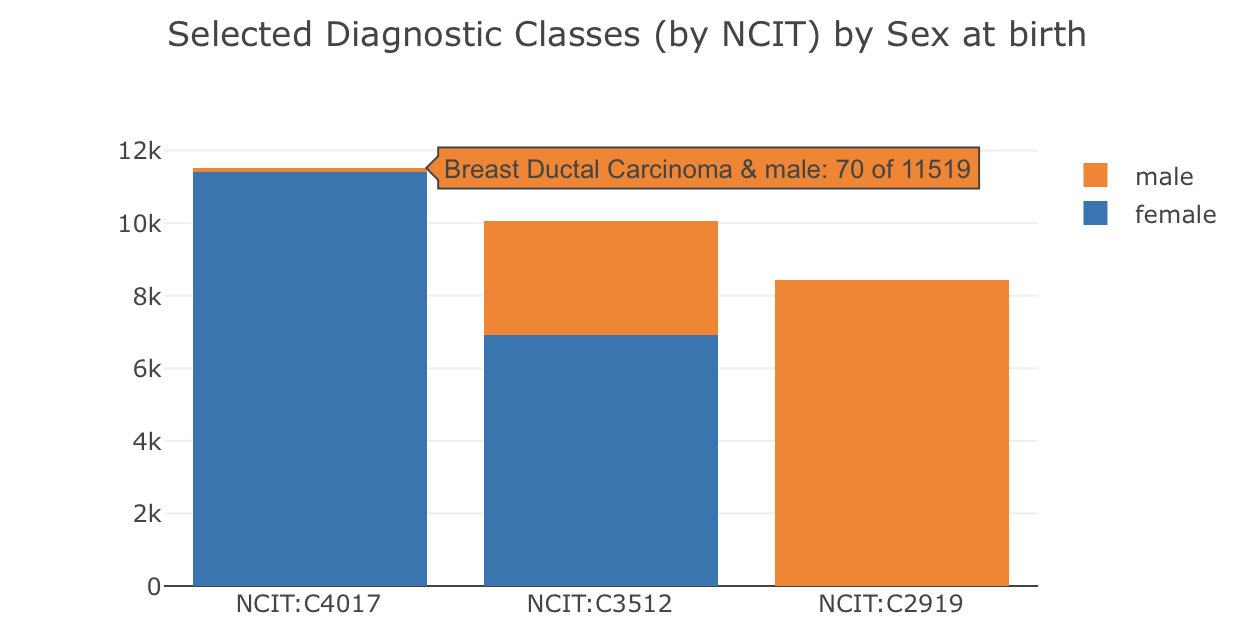
- there are now 2 concepts in the
conceptslist and theconceptValuesin the distribution are observed combinations of values for both concepts, in the same order - the
countindicates the number of times this combination was observed (e.g. 8421 cases of "male" & "Prostate Adenocarcinoma" but 0 cases for "female" & "Prostate Adenocarcinoma")
The stacked bar chart was generated in Plotly.js from the Beacon 2D aggregation in the example below, directly derived from the response JSON on the Progenetix site and reflecting the resource's content.
{
"scope": "individual",
"concepts": [
{"id": "selectedDiseases", "label": "Selected carcinoma types"},
{"id": "sexAtBirth", "label": "Sex at birth"}
],
"distribution": [
{
"conceptValues": [
{"id": "NCIT:C2919", "label": "Prostate Adenocarcinoma"},
{"id": "NCIT:C20197", "label": "male"}
],
"count": 8421
},
{
"conceptValues": [
{"id": "NCIT:C2919", "label": "Prostate Adenocarcinoma"},
{"id": "NCIT:C16576", "label": "female"}
],
"count": 0
},
{
"conceptValues": [
{"id": "NCIT:C4017", "label": "Breast Ductal Carcinoma"},
{"id": "NCIT:C16576", "label": "female"}
],
"count": 11449
},
{
"conceptValues": [
{"id": "NCIT:C4017", "label": "Breast Ductal Carcinoma"},
{"id": "NCIT:C20197", "label": "male"}
],
"count": 70
},
{
"conceptValues": [
{"id": "NCIT:C3512", "label": "Lung Adenocarcinoma"},
{"id": "NCIT:C16576", "label": "female"}
],
"count": 6928
},
{
"conceptValues": [
{"id": "NCIT:C3512", "label": "Lung Adenocarcinoma"},
{"id": "NCIT:C20197", "label": "male"}
],
"count": 3112
}
]
}
Definition of Aggregation Concepts¶
Similar to .../filtering_terms/ (see filters) beacons should indicate their
supported aggregations in the .../aggregation_terms/ endpoint which allows clients
to dynamically discover the available aggregations and their semantics. Aggregation
terms provide single, i.e. 1-dimensional concepts, usually referring to a
single property in the data model. 2D aggregations - reporting the occurrence of
intersecting values, e.g. combining concepts for "diseases" and "sex at birth",
are simply derived from those.
Aggregation concepts can have additional modifiers:
filters- to limit the aggregations to subsets of the data, e.g. to a selection of disease codes- If
filtersare indicated for an aggregation concept, only aggregations for the concept's property fulfilling the individual filters will be reported - Without
filtersall values for the property will be reported (with potential limits imposed by the beacon). - The usual filter definitions apply; i.e. if an ontology term is used as
filter value the count will include all records with this term or any of
its child terms (unless an
includeDescendantTermsflag is set toFalse)
- If
-
splits- to partition continuous data such as age values, followup times or other numeric measurements into countable bins- Note:
splitsare upper & exclusive boundaries of bins, following established practices (cf.$splituse in MongoDB aggregation pipelines)
- Note:
-
a
sortedflag, to indicate that the aggregation results are returned in a pre-sorted order with some inherent meaning (e.g. age bins) - a
formatproperty to indicate the format of the values, e.g. for age splits provided in ISO8601 duration format
Basic example¶
id: sampleOriginDetails
label: Anatomical Origin
description: >-
Counts for anatomical sites in matched biosamples
property: biosample.sample_origin_detail.id
filters Example¶
selectedDiseases:
id: selectedDiseases
label: Selected Diagnostic Classes (by NCIT)
property: individual.diseases.disease_code.id
filters:
- id: NCIT:C2919
label: Prostate Adenocarcinoma
- id: NCIT:C4017
label: Breast Ductal Carcinoma
- id: NCIT:C3512
label: Lung Adenocarcinoma
splits Example¶
ageAtSampleCollection:
id: ageAtSampleCollection
label: Age at sample collection
description: >-
Age at diagnosis (sample collection...)
property: biosample.collection_moment
format: iso8601duration
splits:
- value: P18Y
label: < 18 years
- value: P65Y
label: < 65 years
- value: P120Y
label: 65+ years
sorted: True
Requesting Aggregations¶
Use aggregated Granularity
Aggregation responses are invoked by setting the granularity parameter to aggregated in the request: ?requestedGranularity=aggregated. This indicates that the client is not interested in record level responses but rather in aggregated summaries of the data content.
To request specific aggregations from the ones indicated at the .../aggregation_terms
endpoint clients can use the aggregators query parameter which itself is an array
of arrays of concepts.
In this example 2 aggregations are requested: A simple 1D aggregation for the sampleOriginDetails concept and a 2D aggregation for the combination of selectedDiseases and sexAtBirth.
"aggregators": [
[
{"id": "sampleOriginDetails"}
],
[
{"id": "selectedDiseases"},
{"id": "sexAtBirth"}
]
]
The non normative GET example uses a standard comma concatenation for
the outer aggregators list and square brackets [] for nesting and indication
of intersecting concepts.
?aggregators=[sampleOriginDetails],[selectedDiseases,sexAtBirth]
